  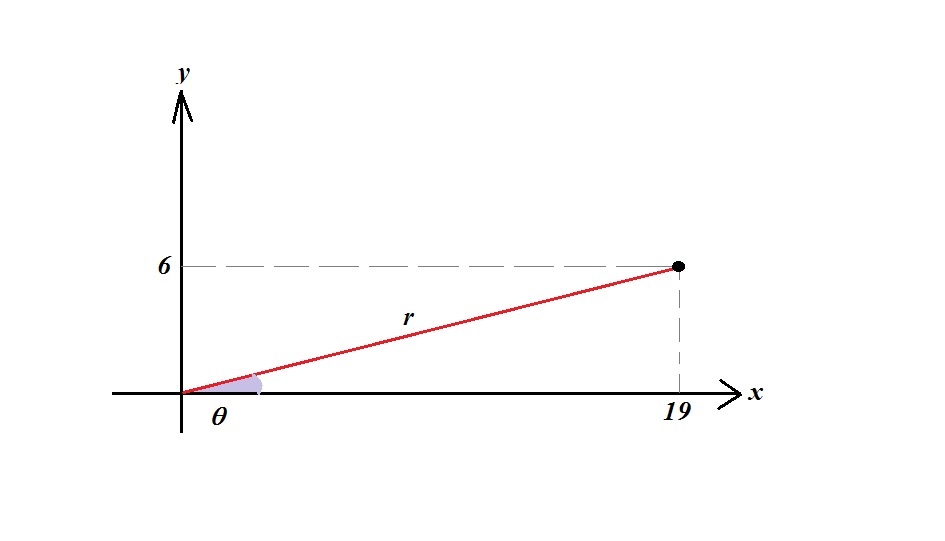
If you don't want to use one of the above free tools, and don't even want to look at the how it was done in the free Python code I mentioned, then still the procedure is doable (though a bit tedious).ĭistances: Calculate the Euclidean distances between each pair of coordinates in your molecule. If for some reason you want to code this in Python yourself: There's many such tools available for free online, including this Python program. A python code to do this (requires downloading the code): I have done it for you and provided the ZMAT file at the bottom of this answer. How to do this, has been asked here on the Chemistry Stack Exchange, and my answer was to use this free tool in which you just copy and paste the 3 spatial coordinates along with the name of the element, for each row of the XYZ file. 
What you are describing, is the conversion from XYZ format to ZMAT format. 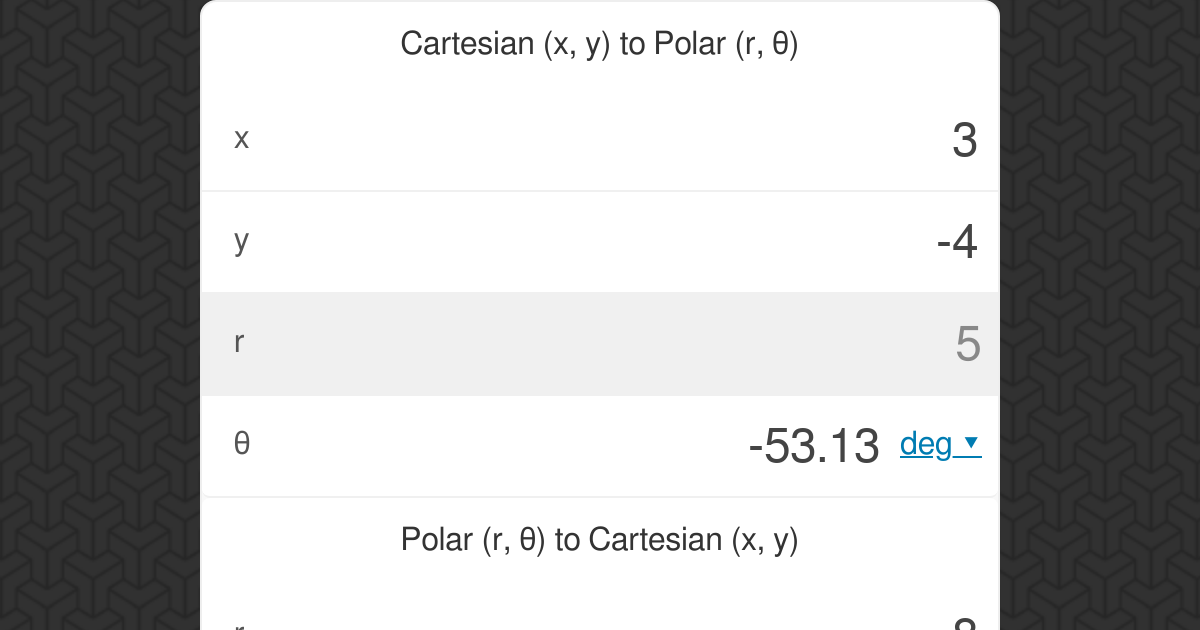
Here's an example script inspired by rsdel's answer which uses ASE: import ase.ioĪtom = ase.io.read(io.StringIO(xyz_file), format = 'xyz')Įasiest free tool to do this (no download necessary): The ASE library has an atom object with built-in get_angle, get_dihedral and get_distance methods that do just that. From the xyz parameters, I need to find the distance between atoms, angle and dihedral between atoms. I want to make a python script that will load an xyz file. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
